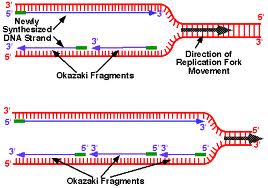
Any protein which can influence either the dinucleotide synthesis rate or primase-ssDNA binding affinity will also play a key role in the regulation of primer synthesis initiation. The rate-determining step is the first phosphodiester bond formed. Using artificial ssDNA templates, primase has been shown to be the slowest and most error-prone polymerase yet studied. This trinucleotide initiation specificity has been shown to be an intrinsic property of pure primase in vitro. 3 Overhang of Adaptor 1 ensured that it could only be ligated to Okazaki fragments’ 3 ends and 5 overhang of Adaptor 2 made it only ligated to Okazaki fragments’ 5 ends. Important among these is that Okazaki fragments are initiated in vivo from d(CTG) most of the time. The strategy for the library construction of Okazaki fragments was basically the same as previously described with minor revision 17 (Fig. Many of the observations concerning primer synthesis initiation in vivo have been reproduced by several of the in vitro assay systems. Primase has ¿RNAP¿ motif, a sequence found in all RNA polymerases which may be involved in chain initiation. The function of the zinc finger may be to bind ssDNA in a sequence-specific manner. The region with the putatuve zinc binding motif is the most highly conserved portion, including more than 25% of identical residues among bacterial primases. This motif can be considered to be involved in phosphodiester bond formation. The strand is synthesized in short segments, named Okazaki fragments, after their discoverer (Sakabe and Okazaki 1966 Okazaki et al. (2) named these small fragments Okazaki fragments, as they have been called since. One of the magnesium binding motifs resembles the ¿active magnesium¿ motif found in all DNA and RNA polymerases. A review of the current state of molecular biology published that year. Using amino acid sequence analysis the structure of Escherichia coli primase responsible for binding zinc, at least three magnesium, and DnaB helicase has been identified. As a primase, it has structural and functional features for initiating chain synthesis on ssDNA. Pol elongates Okazaki fragments, which are 150 nt in length, and also matures these fragments. After a switch to Pol, the enzyme can correct replication errors that were made by Pol during initiation. As RNA polymerase, it has structural and functional features for initiating chain synthesis. Each Okazaki fragment on the lagging strand is initiated by the primase activity of Pol Primase. In common with all DNA and RNA polymerases, primase has structural and functional features involved in polymer elongation. Primase is the ssDNA-dependent RNA polymerase that synthesizes RNA primers during DNA replication.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
